What’s new in this update?
- The reference list of available assays has been updated.
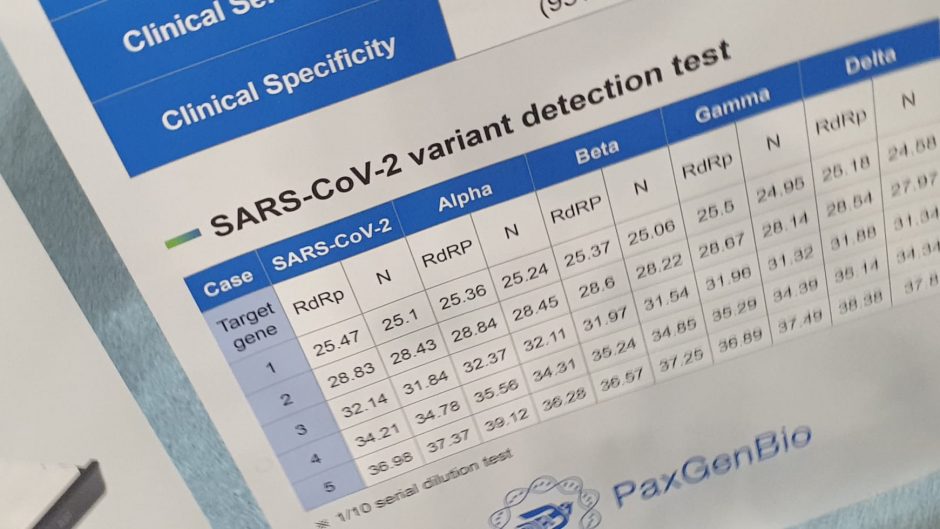
- Delta and Omicron variant assays have been included.
- The Rapid Antigen Detection Tests chapter has been updated to include information related to the Health Security Committee’s list of mutually recognized tests in EU/EEA countries.
Key messages
- Several SARS-CoV-2 VOCs have emerged in recent months and monitoring their circulation in all countries is of vital importance to prevent and control the spread of VOCs.
- Complete sequencing of the SARS-CoV-2 genome, or at least complete or partial sequencing of the S gene, should be used to confirm infection with a specific variant and to characterize the variant.
- For early detection and estimation of VOC prevalence (or where sequencing capacity is limited), alternative methods such as NAAT-based diagnostic tests should be used. To contain or delay the introduction of a VOC, ideally, positive samples should be tested using a NAAT-based assay that can offer the advantage of rapid results.
- When NAAT-based methods are employed, sequencing should be used to characterize at least a subset of the variants.
- In case of the low prevalence of a VOC in the population and when the goal is to delay the introduction and spread of the VOC, ideally, all NAAT-based results indicative of the VOC should be confirmed by sequencing.
- The selection of samples and methods is key and will depend on the objectives (eg, evaluating the circulation of the different variants of SARS-CoV-2 using representative samples of the community, or genomic characterization to monitor the evolution of the virus and inform decisions about vaccine composition or outbreak analyses).
- Assay validation should be performed to ensure that the laboratory testing system is working properly for circulating viruses.
- Laboratories must remain vigilant to ensure that they detect reduced sensitivity or failure to detect/identify circulating variants by different PCR or antigen-based assays.
- If diagnostic capacity is insufficient, priority should be given to severe cases, fatal cases, and cases with suspected high contagiousness of the pathogen that caused the outbreak, especially among those vaccinated or with a history of COVID-19.
- SARS-CoV-2 consensus sequences should be submitted to GISAID or other public databases. Raw sequences should also be submitted to the European Nucleotide Archive (ENA).
- Detection of new VOCs or outbreaks of currently circulating VOCs must be reported immediately through the Early Warning and Response System (EWRS), while variant detections must be reported to Tessy on a weekly basis.
SARS-CoV-2 variants with CDC classification
1. Lineage (VOC).
Originally from the UK, The B.1.1.7 variant contains many genetic mutations. the variant was first identified in September 2020. Since then, the variant it can been found in numerous countries around the world. The tension was found in the United States in late December 2020. It has become the most common strain in the US and is associated with higher transmissibility (i.e., more efficient with rapid transmission) and increased mortality.
2. Lineage B.1.351 (VOC).
In South Africa, the variant B.1.351 arose independently of B.1.1.7, although the variant shares some genetic mutations with the strain. B.1.351 was the first identified in South Africa in samples dating from early October 2020. Since then, cases have been detected in several countries, including the US in 2021.
3. Lineage P.1 (VOC).
The P.1 variant was first identified in Brazil Travellers. The variant has 17 unique genetic mutations, including three in the receptor-binding domain (RBD) region of the spike protein. Surveillance detected the variant in the US in late January 2021. Variant P.1 represents a branch of the B.1.1.28 lineage. Evidence suggests that some of the mutations in the P.1 variant may affect transmissibility and antigenic profile. The appearance of the variant and the association with higher viral density raised concerns about possible increased transmission or a propensity for reinfection.
4. B.1.427 and B.1.429 (VOC).
Both strains of coronavirus California are now officially characterized. Deformations can be 20% more transmissible than the common strains found in California, Nevada, Arizona, Wisconsin and Virginia.
5. B.1.526 (VOI).
First detected in New York in November 2020, the B.1.526 is a variant of interest. For February 2021 mutations appeared in about 25% of the entire COVID genomes.